🤝 Top Collaborators
🤓 Latest Submissions
DemoPilot - Autonomous Demo Video Agent
Every software team needs demo videos, for investors, judges, launches, and onboarding. Today, creating one means recording in OBS or Loom, re-recording every mistake, editing in timeline software, and hiring voiceover artists or settling for robotic TTS. A 3-minute demo can take min 2–4 hours. Who has time? Introducing DemoPilot, a fully autonomous AI agent that generates production-ready demo videos from a URL. The pipeline is end-to-end, with no human touches required. It starts with Capture, where Playwright visits the target URL and captures a screenshot plus page structure, including nav links, buttons, headings, and form fields. Next is Script, where Gemini 3 Flash analyzes the screenshot and elements, then writes a narrated demo script with actions like click, type, scroll, navigate, and hover using real CSS selectors. In Record, Playwright drives headless Chromium at 1920x1080, adding gold click glows, smooth scrolling, and character-by-character typing for a natural feel. Narrate uses edge-tts to generate neural voiceover with 10+ Azure voice options, timed to each segment. Finally, Compose uses ffmpeg to merge recording and narration into a polished H.264 MP4. DemoPilot also supports voice-to-script: users speak their demo intent into a mic, Speechmatics transcribes it, and Gemini turns it into a structured script. AI Quality Assurance uses Gemini Vision to review six final-video frames, score quality, and flag issues. Other features include cookie consent auto-dismissal for 80+ CMP selectors like OneTrust, Ketch, and Cookiebot, deterministic output for CI/CD workflows, and full manual override so users can edit every segment before rendering. DemoPilot reduces demo video production from hours to minutes. It is built for hackathons, SaaS companies, and developer advocates who need professional demos without professional video editors. Both landing page showcase videos, that were generated by DemoPilot, from URL to finished MP4, untouched by human hands.
19 May 2026

Meridian: Autonomous LC Intelligence
Meridian is an autonomous Letter of Credit intelligence platform that transforms global trade finance. Letters of Credit govern $2.3 trillion in annual cross-border trade across 175 countries under the UCP 600 framework (WTO), yet LC issuance takes 5–21 days of manual bank review with a 60–80% first-presentation rejection rate (ICC Banking Commission). A single shipment involves up to 50 documents exchanged between 30 stakeholders, only 1–2% handled digitally. Meridian solves this with 9 AI agents orchestrated via LangGraph and powered by Gemini 3 Flash. Three intelligence agents analyze applicant documents in parallel — Importer Intelligence for credit scoring and financials, Exporter Intelligence for counterparty risk and export history, and Trade Intelligence for sanctions screening and HS code validation. Their outputs feed into Risk Synthesis for unified risk scoring, UCP 600 Compliance which checks 27 articles plus 6 eUCP v2.1, and LC Generator — which produces a human-readable LC document, a production-ready SWIFT MT700 banking message, and an Officer Decision Brief. The entire issuance pipeline completes in approximately 90 seconds. Three additional examination agents then verify presentation documents field-by-field against issued LC terms, catching discrepancies like HS code misclassifications (8457.10 vs 8459.61) and insurance under-coverage (UCP 600 Article 28) before the bank ever sees them. Gemini's 1M token context window is architecturally essential — all documents are analyzed in a single pass with no chunking, enabling cross-document synthesis that catches issues spanning multiple documents simultaneously. $9.7T addressable market, $2.5T unmet trade finance gap (ADB 2025), zero AI competitors in LC intelligence.
19 May 2026
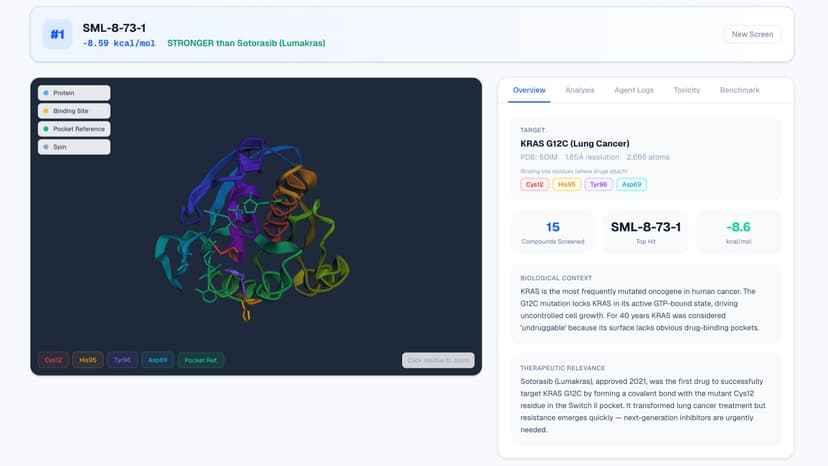
CatalystMD: AI-Powered Drug Discovery Platform
CatalystMD automates early drug discovery by screening real drug compounds against disease protein targets such as COVID-19 protease, HIV-1 protease, EGFR lung cancer, and KRAS G12C (once considered undruggable for 40 years) using real computational chemistry. Team of 5 AI agents work in sequence, each passing results to the next: 1. Target Analyst Agent: Downloads the real 3D protein structure from the Protein Data Bank, identifies the binding pocket where drugs attach, and analyzes the biological context using Qwen 2.5-7B. 2. Molecular Dynamics Engine: Docks each compound into the binding pocket using AutoDock Vina (physics-based scoring) and runs energy minimization with OpenMM on the AMD MI300X GPU using the AMBER14 force field. 3. Binding Scorer Agent: Ranks all candidates by binding affinity and compares each one to the current FDA-approved drug for that target. Qwen generates a scientific interpretation of the results. 4. Toxicity Screener Agent: Checks toxicity & drug-likeness using Lipinski's Rule of Five and screens for PAINS (Pan-Assay Interference Compounds) using RDKit. 5. Discovery Reporter Agent: Generates a complete scientific brief with structural analysis, rankings, toxicity data, and recommended next steps. Targets tested: COVID-19 Main Protease (6LU7), KRAS G12C Lung Cancer (6OIM), EGFR Kinase Lung Cancer (1M17), HIV-1 Protease (1HIV) Real computation, not mock data: CatalystMD uses real X-ray protein structures from the RCSB Protein Data Bank, real compound SMILES from published literature, AutoDock Vina physics-based docking scores, OpenMM energy minimization, RDKit molecular property calculations, and Qwen 2.5-7B analysis served through vLLM. The AMD MI300X powers both the OpenMM molecular simulation through ROCm OpenCL and AI inference through vLLM. Tech stack: AMD MI300X, AutoDock Vina, Meeko, Open Babel, OpenMM, AMBER14, Qwen 2.5-7B-Instruct, vLLM, LangGraph, RDKit, PDBFixer, FastAPI, Python 3.12, Next.js 16, and 3Dmol.js.
10 May 2026
.png&w=256&q=75)
.png&w=640&q=75)




